head(cars) speed dist
1 4 2
2 4 10
3 7 4
4 7 22
5 8 16
6 9 10There are lots of ways to make plots in R. These include so-called “base R” (like the plot()) and on packages like ggplot2.
Let’s make the sampe plot with these two graphics systems. We can use the inbuilt cars dataset:
shortcut: option+command+i(insert)
head(cars) speed dist
1 4 2
2 4 10
3 7 4
4 7 22
5 8 16
6 9 10With “base R” we can simply
plot(cars)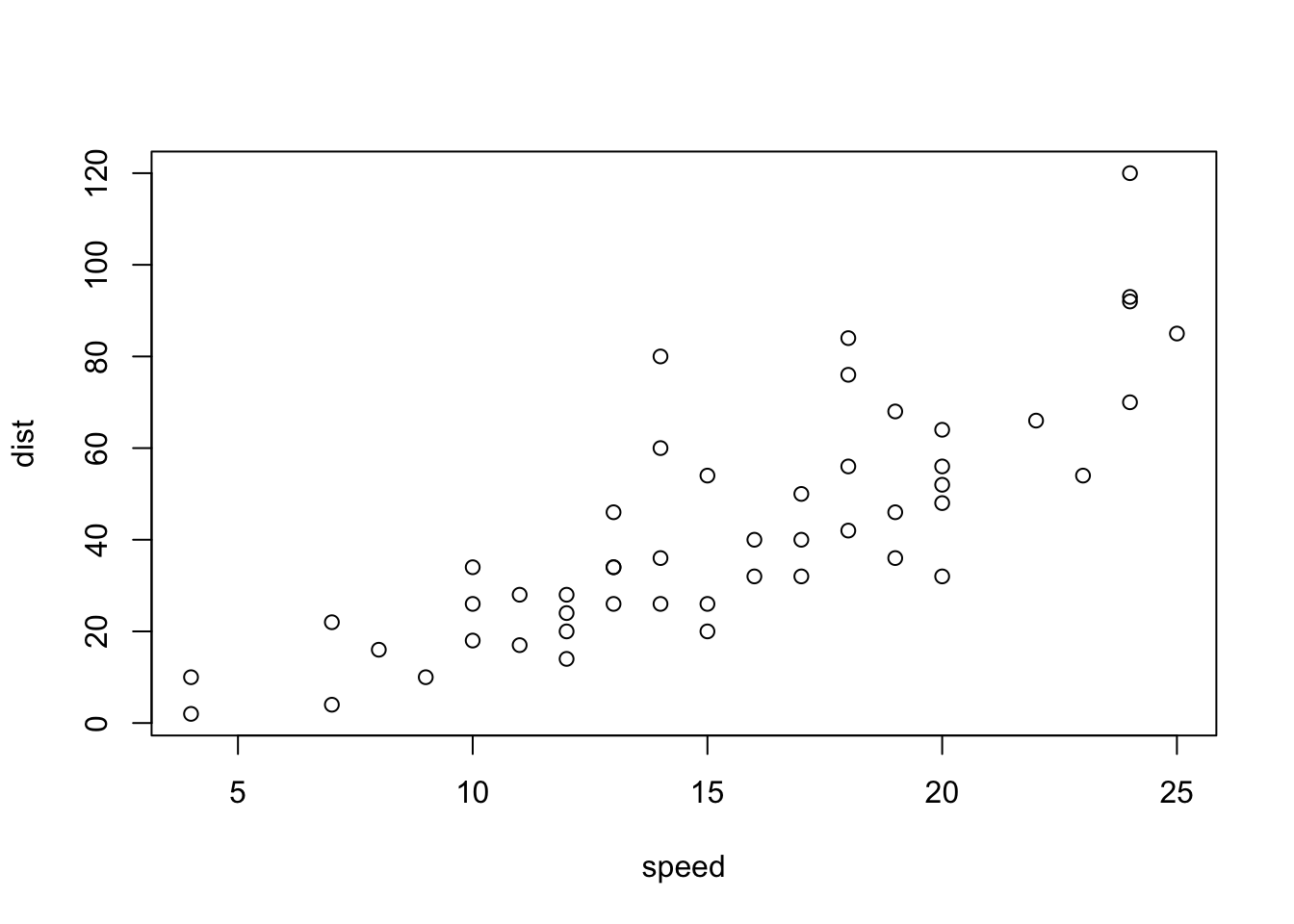
Now let’s try ggplot. First I need to install the package using install.packages("ggplot2").
N.B. We never run an
install.packages()in a code chunl otherwise we will reinstall needlessly every time we render the document.
#———-not use for now————-
url <- "https://bioboot.github.io/bimm143_S20/class-material/up_down_expression.txt"
genes <- read.delim(url)
head(genes) Gene Condition1 Condition2 State
1 A4GNT -3.6808610 -3.4401355 unchanging
2 AAAS 4.5479580 4.3864126 unchanging
3 AASDH 3.7190695 3.4787276 unchanging
4 AATF 5.0784720 5.0151916 unchanging
5 AATK 0.4711421 0.5598642 unchanging
6 AB015752.4 -3.6808610 -3.5921390 unchangingnrow(genes)[1] 5196ncol(genes)[1] 4table(genes$State)
down unchanging up
72 4997 127 table(genes$State)/sum(table(genes$State))
down unchanging up
0.01385681 0.96170131 0.02444188 1+1[1] 2